Deep Learning for Patch Clamp Electrophysiology
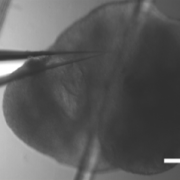
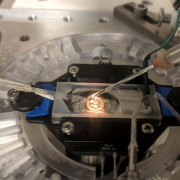
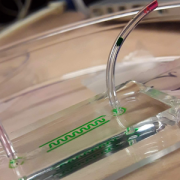
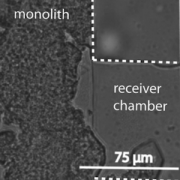
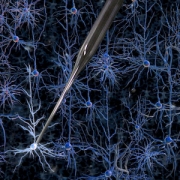
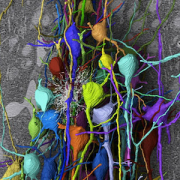
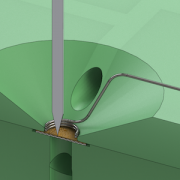
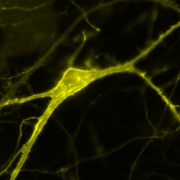
With the recent advances in deep learning enabling rapid and accurate identification of complex structures, it’s application to in vitro patch-clamp electrophysiology was inevitable. We are exploring the application of these techniques to enhance the capabilities of our fully-automatic patch clamp robots. For example, we have demonstrated fully automatic, real-time detection of healthy neurons within traditional DIC images and vision-based techniques to correct for hardware error stack-up during pipette localization.